Welcome!
This is the personal homepage of Lazaros Gallos.
I am an Associate Director and Research Professor at DIMACS at Rutgers University, where I am also directing the long-running DIMACS REU summer program. DIMACS fosters research and educational programs on topics that lie at the interface of discrete mathematics and theoretical computer science.
I am a member of the Editorial Boards at Scientific Reports, at PLOS ONE, and at Entropy. I have also been an Associate Editor for the highly selective APS journal Physical Review X.
In the past, I was a Research Associate at the Department of Ecology of the
Rutgers University, working with Prof. Nina Fefferman.
Before that, I was working with Prof. Hernan Makse at the City College of New York.
I received my PhD in Computational Physics from the Physics Dept. in Univ. of Thessaloniki (advisor Prof. Panos Argyrakis).
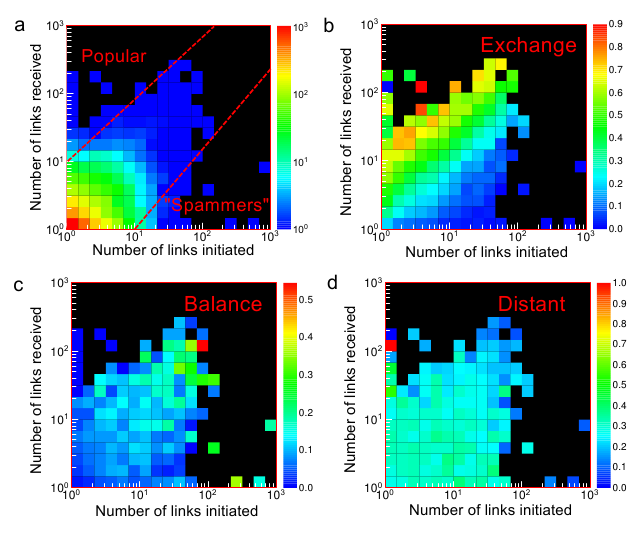
My research interests are broad and cover a lot of inter-disciplinary ground. For the last few years I have been working on complex networks science. Currently, I work on (a) understanding fundamental mechanisms behind online social interactions,
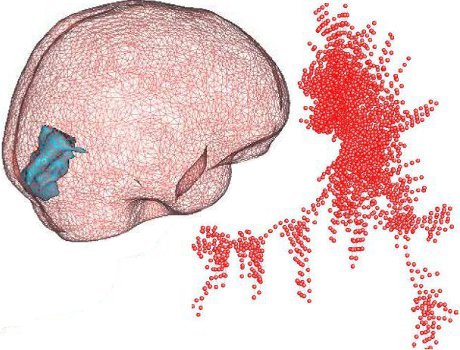
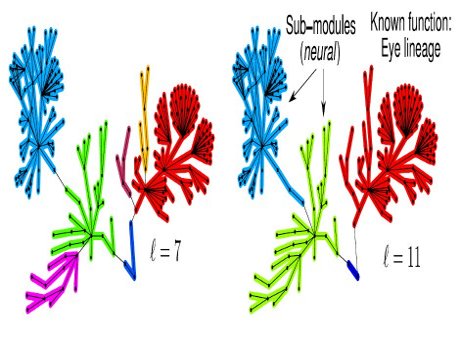
(b) epidemics spreading, (c) modules organization in the brain, and (d) applications of Machine Learning in Network Science.






Research Highlights
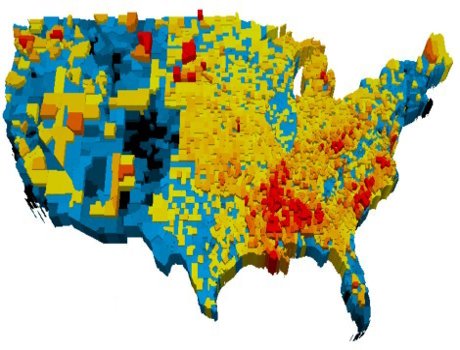
Network comparison
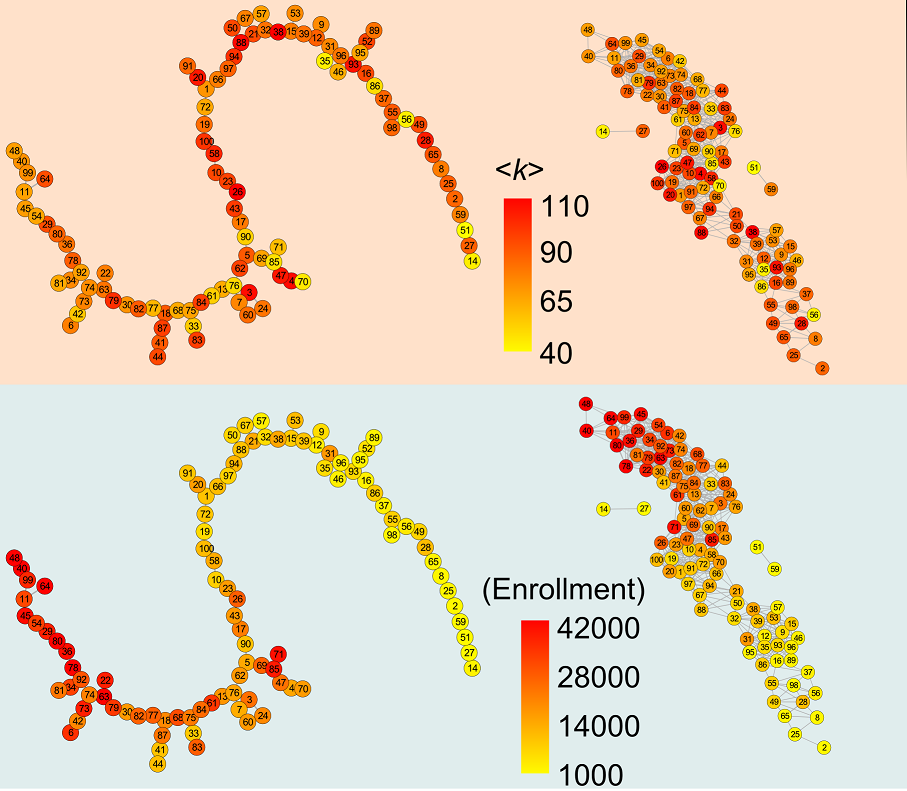
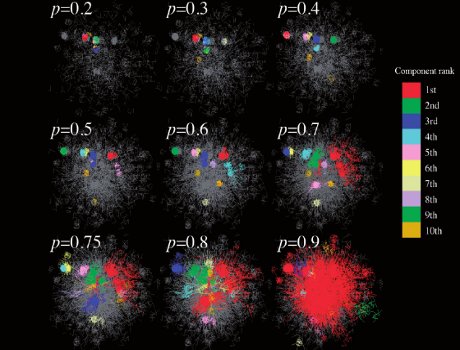
There are many practical applications where we want to compare two structures, such as complex networks. We proposed a method where we sample two networks at a given scale and compare the link density distribution at this scale. The method works very well, and can be used to identify effective classifiers.
To compare two networks, you can download the source code ntangle.c. Check also the README file.Connections aren't everything

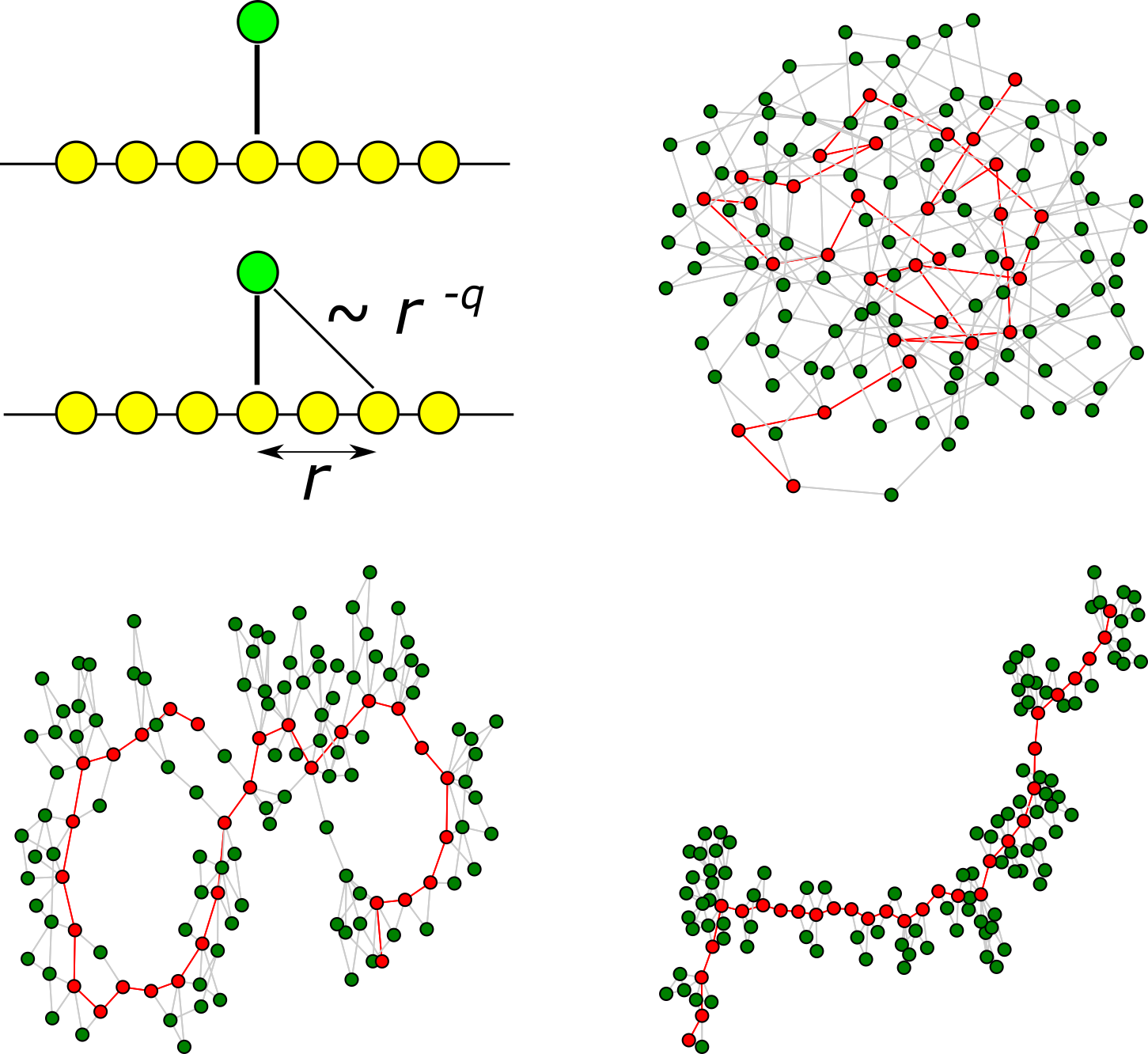
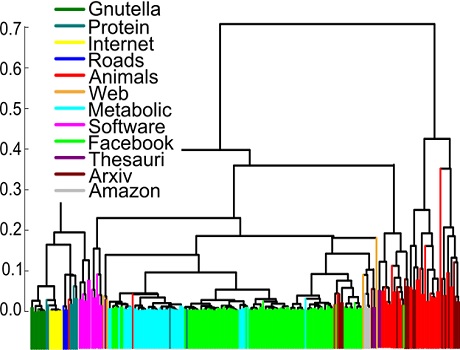
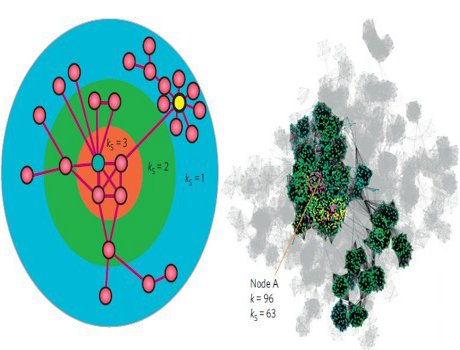
You want to spread a message in a social network. What would be your choice of the starting point for more efficient spreading? Surprisingly, it is not always the most connected individual; location can be more important than the number of connections.